17April, 2018
Tricking hyperplanes
Tricking hyperplanes
## Setting default kernel parameters
Support Vector Machines
Instead of requiring a clean separation, penalize misclassified points by how far they are misclassified.
Label two classes by \(y_i=\pm1\). The separating hyperplane earlier is the minimizing \(w, b\) of \[ \min \|w\| \qquad\text{subject to}\qquad y_i(w\cdot x_i-b) \geq 1 \quad\forall i \]
If there is no separating hyperplane, the conditions cannot be fulfilled. Instead weaken the condition, using a loss function that penalizes the misclassification. \[ \min_{w,b}\left[ \frac1n\sum\max(0, 1-y_i(w\cdot x_i-b)) +\lambda\|w\|^2 \right] \]
Support Vector Machines
SVMs work with the optimization target \[ \min_{w,b}\left[ \frac1n\sum\max(0, 1-y_i(w\cdot x_i-b)) +\lambda\|w\|^2 \right] \]
If \(y_i\) is confidently correctly classified, the penalty \(\max(0,1-y_i(w\cdot x_i-b))\) is 0. When in doubt, the penalty increases, and when misclassified the penalty will exceed 1. The \(\|w\|^2\) term narrows the margin.
Support Vector Machines
SVMs work with the optimization target \[ \min_{w,b}\left[ \frac1n\sum \color{green}{\max(0}, 1-y_i(\color{orange}{w\cdot x_i-b})) \color{purple}{+\lambda\|w\|^2} \right] \]
- Ignore correctly classified points
- Linear decision boundary
- Maximize margin
Since correctly classified points are ignored in the optimization, only a few vectors contribute to determining \(w, b\).
These are called the support vectors of the decision boundary.
Support Vector Machines
Support Vector Machines
Return of the kernel
Where Support Vector Machines really gain power is in using kernel methods: it can be difficult to even find a library that does not immediately use kernels.
The Kernel Trick replaces \(w\cdot x\) with \(\phi(w)\cdot\phi(x)\) for some \(\phi\) mapping vectors to some higher (possibly infinite) dimensional space.
The key is that \(\phi(x)\) never is needed: only \(K(w,x)=\phi(w)\cdot\phi(x)\) is ever used.
Common kernels
| Formula \(K(x,y)=\) | Name in R::kernlab |
Name in Python.scikit-learn |
|---|---|---|
| \(x\cdot y\) | vanilladot |
linear |
| \((\gamma (x\cdot y)+r)^d\) | polydot |
poly |
| \(\tanh(\gamma(x\cdot y)+r)\) | tanhdot |
sigmoid |
| \(\exp[-\gamma\|x-y\|^2]\) | rbfdot |
rbf |
| \(\exp[-\gamma\|x-y\|]\) | laplacedot |
R::kernlab also provides besseldot (Bessel function on \(x\cdot y\)), anovadot (product of RBF kernels), splinedot (cubic spline) and stringdot (string kernels)
Kernel SVC
Kernel SVC
Kernel SVC
Dialects of Support Vector Classifiers
C-SVC Use an equivalent reformulation producing a parameter \(C\) controlling emphasis on keeping the no point's land region clear.
NuSVC Introduce a parameter \(\nu\in(0,1]\) producing an upper bound on the fraction of training errors and lower bound on the fraction of support vectors.
In R
The package kernlab has a good SVM implementation
model = kernlab::ksvm(class ~ predictor1 + predictor2, data=data, kernel="rbfdot")
Also takes type to change between classification (C-svc, nu-svc, …), regression (eps-svr, nu-svr, …) and novelty detection (one-svc) and kpar to provide details (\(\gamma\), \(d\), \(r\), …) for the kernels.
Use as
classes = predict(model, data) class.probs = predict(model, data, 'probabilities')
In Python
In scikit-learn the module svm contains support vector machines of various versions
| Model | Type |
|---|---|
svm.LinearSVC |
Linear support vector classification (optimized) |
svm.NuSVC |
Nu-support vector classification |
svm.SVC |
C-support vector classification |
Cautions and best practice
SVMs are not scale invariant: whiten or rescale each attribute to the same range ([0,1] or [-1,1] common choices) data to produce a well-performing method.
This is done automatically in kernlab, and with preprocessing pipelines in scikit-learn.
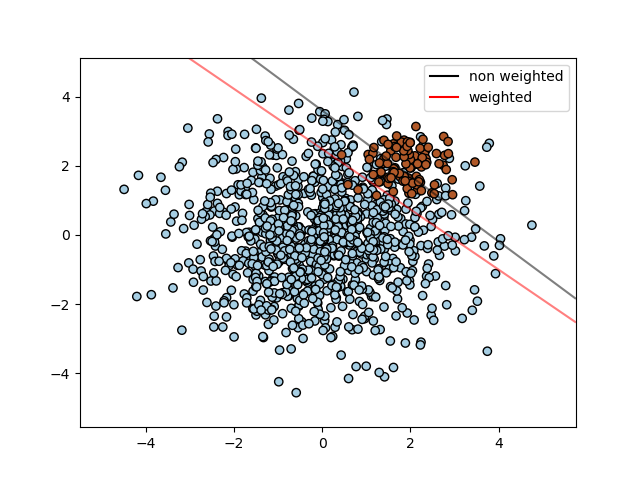
SVCs are sensitive to unbalanced data. By weighting data points differently, this can be compensated for. Command option to calculate weights in scikit-learn, and class weights can be produced by hand in kernlab.
Unbalanced data
Trees and Forests
Decision tree
One way to produce a classification is to split data space by one coordinate value at a time, creating as homogenous as possible regions.
Splits should be chosen to minimize impurity: maximize agreement within the parts. For region \(m\) and class \(k\), write \(\hat p_{mk}\) for the proportion of class \(k\) in region \(m\).
- Misclassification error: \(1-\hat p_{mk}\)
- Cross-entropy: \(-\sum\hat p_{mk}\log\hat p_{mk}\)
- Gini index: p_{mk}(1-p_{mk})$
Decision tree
Bagging: Bootstrap aggregation
- Bootstrap: sample \(N\) times with replacement from the \(N\) data points, average across samples
- Aggregation: average results from bootstrap samples.
If each of \(B\) bootstrap draws has variance \(\sigma^2\), but correlation \(\rho\), then \[ \mathbb V = \rho\sigma^2+\frac{1-\rho}{B}\sigma^2 \]
The second term decays with \(B\). The first term remains: bagging with decorrelated predictions improves performance.
Random Forest algorithm
- For b from 1 to B
- Draw a bootstrap sample \(Z^*\) of size \(N\) from the training data
- Grow a classifier tree \(T_b\) repeating until specified tree size:
- Select \(m\) variables at random
- Pick the best split-point among the \(m\)
- Split the node into two daughter nodes
- Return collection \(\{T_b\}\) of trees.
To predict, run all \(B\) trees and pick majority vote for predicted class.
The random selection of variables improves decorrelation and reduces the classifier variance. This is specific to very non-linear methods – linear predictors get very little decorrelation.
Example
Single-pass cross-validation
An out-of-sample error estimate can be built while training a random forest:
For each \(z_i=(x_i,y_i)\) construct the predictor by voting from only the trees built on bootstrap samples not containing \(z_i\).
Monitoring this out-of-bag sample errors provides feedback on the number of trees required.
Variable importance
By running out-of-bag samples through predictor trees once unchanged, and then with a single variable permuted among the samples, the impact of any one variable on the classifier can be accumulated.
Boosting
Combines classifiers to reduce bias. Classifiers are combined with weighting – for classifiers \(k_1,\dots,k_j\) the boosted classifier is: \[ C_m(x) = \alpha_1k_1(x) + \dots + \alpha_mk_m(x) \]
At each training step a new classifier is added, and chosen to decrease an error measure.
Boosting
For AdaBoost (Adaptive Boosting), let the total error be exponential loss: \[ E = \sum e^{-y_iC_m(x_i)} \]
Adding \(k_{m+1}\) we would get total error \[ w_i^{(m)} = e^{-y_iC_m(x)} \qquad E = \sum w_i^{(m)}e^{-y_i\alpha_{m+1}k_{m+1}(x_i)} = \\ \sum_{y_i=k_{m+1}(x_i)}w_i^{(m)}e^{-\alpha_m} + \sum_{y_i\neq k_{m+1}(x_i)}w_i^{(m)}e^{\alpha_m} = \\ \sum w_i^{(m)}e^{-\alpha_m} + (e^{\alpha_m}-e^{-\alpha_m}) \sum_{y_i\neq k_{m+1}(x_i)}w_i^{(m)} \]
Boosting
To minimize the total error \[ \sum w_i^{(m)}e^{-\alpha_m} + (e^{\alpha_m}-e^{-\alpha_m}) \sum_{y_i\neq k_{m+1}(x_i)}w_i^{(m)} \] it is enough to minimize \(\sum_{y_i\neq k_{m+1}(x_i)}w_i^{(m)}\). With the minimizing classifier, calculate error rate and weight \[ \epsilon_{m+1} = \sum_{y_i\neq k_{m+1}(x_i)}w_i^{(m)}/\sum w_i^{(m)} \\ \alpha_{m+1} = \frac12\log\left(\frac{1-\epsilon_m}{\epsilon_m}\right) \]
Many generalizations exist, several implemented in the xgboost software package, which performs very well in competitions and applications.
Stacking
Use several classifiers and combine their results with a combiner algorithm. Often, logistic regression is used here.
Summary of ensemble models
| Bagging | Boosting | Stacking |
|---|---|---|
| Random partition | Preference to misclassified samples | Varies |
| Minimize variance | Decrease bias, increase predictive power | Both |
| Combine with (weighted) averages or majority vote | Combine with weighted majority vote | Combine with logistic regression |
| Trained in parallel | Trained in sequence | |
| Good for high variance low bias | Good for low variance high bias | |
| Compensates for overfitting | Compensates for underfitting |
Example
In R: use caret
library(caret) fit = train(Class ~ ., data=training, method="rf")
In R: use caret
| Method | method="..." |
|---|---|
| Classifier tree | rpart, ctree, C5.0, C5.0Tree |
| Random Forest | rf, ranger, rBorist |
| Bagging | AdaBag, treebag, logicBag, bag, |
| Boosting | adaboost, AdaBoost.M1, AdaBag, ada, gamboost, glmboost, LogitBoost, xgbLinear, xgbTree, gbm |
In Python
scikit-learn has the module sklearn.ensemble containing AdaBoostClassifier, BaggingClassifier, GradientBoostingClassifier, RandomForestClassifier.
Also sklearn.tree containing DecisionTreeClassifier and ExtraTreeClassifier.